The fourth most common neurological disorder in the world is epilepsy, according to the Epilepsy Foundation. Individuals who have the disorder experience surges of electrical activity in their brains that can cause recurring seizures. At the base of those electrical signals is a protein called the ionotropic glutamate receptor (iGluR). Problems with this protein cause the seizures, as well as degeneration of nerve cells.
According to Maria Kurnikova from Carnegie Mellon University, when the glutamate receptor “is a little bit too excitable, when it stays open too long, it immediately goes to disorders of the brain.” She would know – she worked with her graduate students and Electrophysiologist Alexander Sobolevsky, an iGluR structure pioneer at Columbia University, to thoroughly examine the motions of iGluR when glutamate attaches to it.
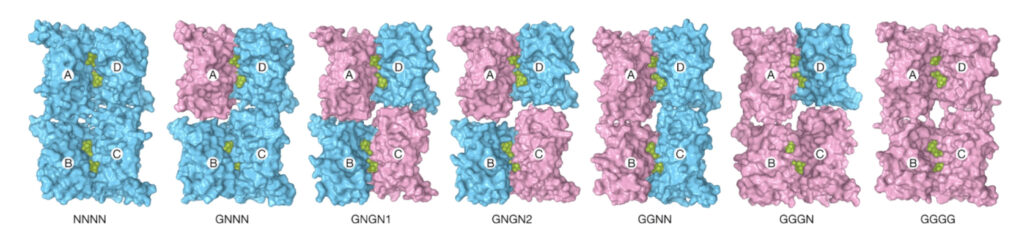
Running molecular dynamic (MD) simulations – among the most complex that Kurnikova’s team had attempted – on Anton 2 and repeating computations using Bridges-2’s powerful GPUs, the scientists found a mixture of kinks with various numbers of glutamate bound. An analysis of the MD data using machine learning artificial intelligence helped tease apart these complex states.
The scientists anticipate additional work to clarify the activity of the glutamate-bound iGluR. But for now, they have shed light on how this important but complicated protein works. It’s the first step in designing new drugs to affect the receptor, giving doctors new options for helping people with problems in this life-critical protein.
There’s one single FDA-approved drug to inhibit [these] receptors – perampanel. But no one knew how it worked on the channel … We produced an enormous amount of simulation data using Anton and [Bridges-2] … It was a difficult problem to simulate the [current] fluctuations, [a] small signal in really noisy data.
Maria Kurnikova, CMU
You can read more about this story here (Published by PSC, Jan. 18, 2023): Anton 2, Bridges-2 Simulations Explain Life-Critical Protein in the Brain
Project Details
Institution: PSC (Pittsburgh Supercomputing Center)
University: Carnegie Mellon University
Funding Agency: This work was supported by the NIH (grant nos. U24 GM129539, U24 GM129541, R01 NS083660, R01 GM128195 and R01 AG065594, R01GM116961, R01 CA206573, R01 NS083660, R01 NS107253 and R01 AR078814). This work also was supported by the NSF (grant nos. 1818213, 1563291, ACI-1548562 and 1818086) and the Simons Foundation (grant no. SF349247).
Grant Number: See above
The science story featured here, allocated through August 31, 2022, was enabled through Extreme Science and Engineering Discovery Environment (XSEDE) and supported by National Science Foundation grant number #1548562. Projects allocated September 1, 2022 and beyond are enabled by the ACCESS program, which is supported by National Science Foundation grants #2138259, #2138286, #2138307, #2137603, and #2138296.


